Gene Mutations Involved in Brain Development and Autism Spectrum Disorders
By: Alexandra Flores Email: flor3662@vandals.uidaho.edu
Home Town: Caldwell, ID High School: Thomas Jefferson Charter School
Major: Biochemistry
Department: Biological Sciences
College: College of Science

Autism Spectrum Disorder (ASD) is a developmental disability characterized by sensory and communication deficits. Recently, diagnosis of ASD has increased, and it is now estimated to affect 1 in 68 children. The symptoms of autism begin to appear during the first three years of life, which coincides with the development of learning ability and a critical period of synaptogenesis. Because synaptic changes are thought to be the basis of all learning, mutations that may affect the neural synapse have been shown to cause the mental retardation phenotype that is associated with autism. Research into the specifics of these synaptic mutations has given evidence that synaptic cell adhesion proteins contribute to autism with at least 28 of the 200 genes associated with autism being identified as cell adhesion molecules (CAMs) or CAM-related.
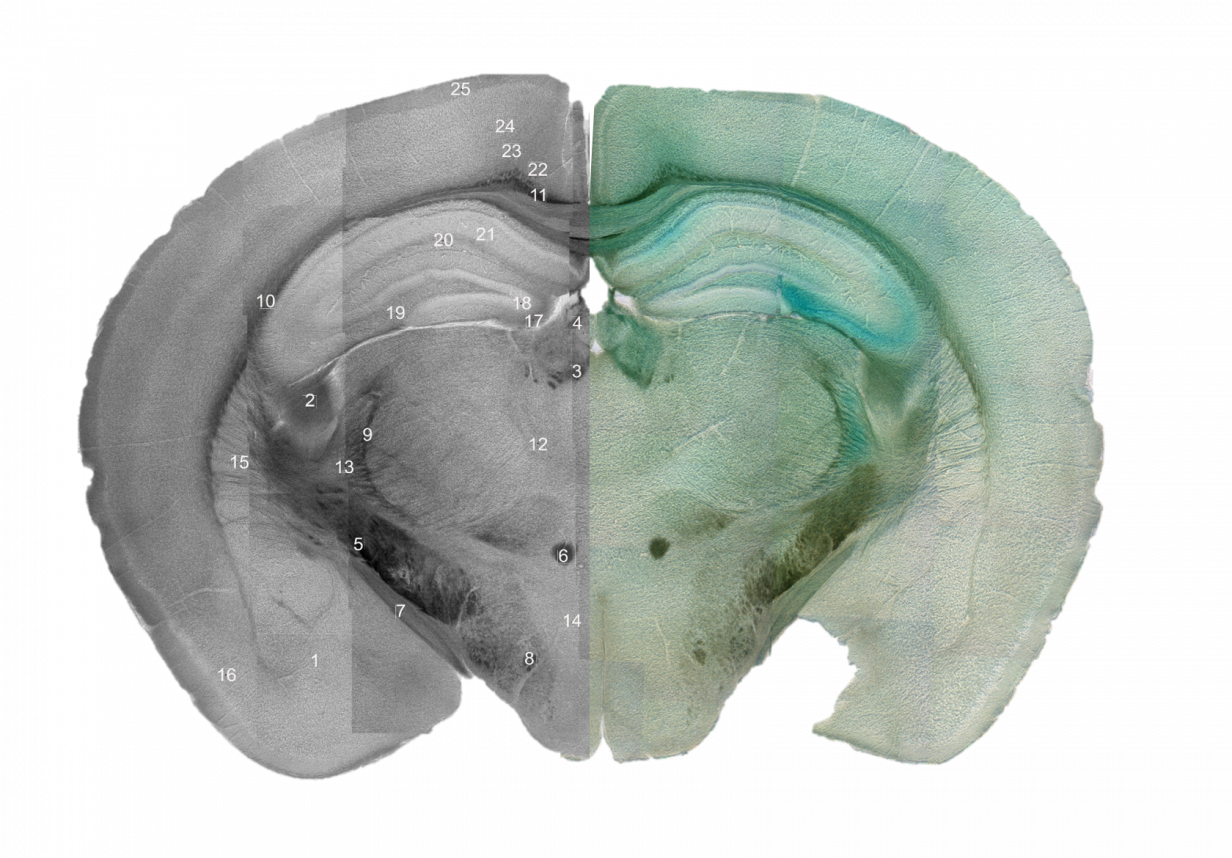
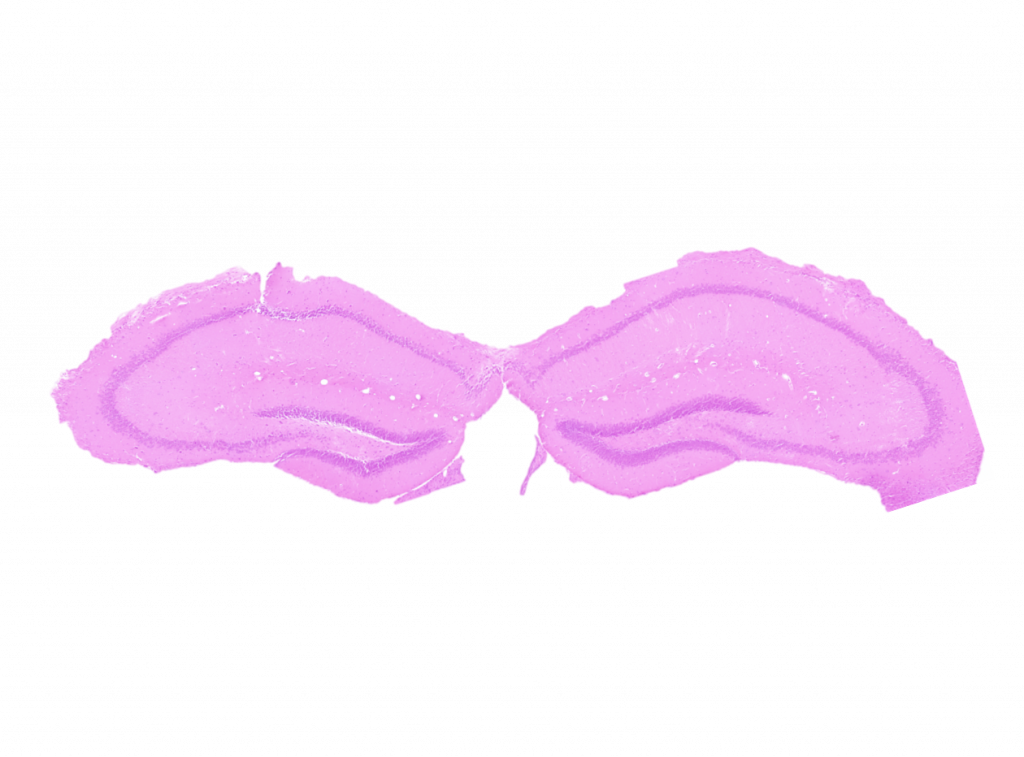
One CAM with expression mutations that have been associated with ASD is Down Syndrome Cell Adhesion Molecule (DSCAM), which functions in neural development, including axon guidance, axonal and dendritic branching and targeting, and synapse maturation. DSCAM has been extensively studied in mice where mutations in the gene have been found to impair aborization, the processes and spacing of a neuron’s dendrites, cell death, axon pathfinding, and synaptic pairing between different types of neurons. However, almost all of these studies have been done only in the retina. Dscam is also expressed in the cerebral cortex, dentate gyrus, thalamus, cerebellum, olfactory bulb, and hippocampal neurons in mice. An analysis of the structure in these major brain regions of Dscam mutant mice can allow for a comparison between the brains of these mice and the brains of people with Dscam expression disorders in order to determine if mice will make a good model to test clinical interventions for ASD treatment. Preliminary results have shown gross defects in Dscam mutant mouse brains, and we aim to analyze the brain regions that are affected in order to better understand DSCAM’s impact on brain function and development.
Genetically-modified mice with loss of function and gain of function phenotypes will be examined to gain a better understanding of DSCAM’s effect on bran function and development. Mouse brains will be analyzed using hematoxylin and eosin staining to identify major brain regions and count cell populations within those regions. The populations will be analyzed according to their organization by comparing cell density. Initial studies will place emphasis on the cerebellum, hippocampus, midbrain, cerebral cortex, and motor cortex. We will also be working closely with computer scientists to generate a digital reconstruction of the major brain regions in order to create a searchable template similar to The Allen Brain Atlas, an online resource that documents different regions of the wild type mouse brain, where the impact of Dscam mutations can be observed.
Products Produced:
| Type: | Title: | Date Published/Presented: | DOI: |
|---|---|---|---|
| Poster | Characterizing Differences in the Superior Colliculus of Genetic Mouse Models for Autism | March 20, 2019 | |
| Poster | Genetic Mutations Involved in Brain Development and Autism Spectrum Disorders | July 31, 2019 | |
| Presentation | Genetic Mutations Involved in Brain Development and Autism Spectrum Disorders | July 31, 2019 | |
| Presentation | Genetic Mutation in the Brain that Contributes to Autism | August 3, 2018 |
Additional Project Information:
Year in College Project Started: Sophomore
Faculty Advisor: Peter Fuerst
Faculty Advisor Email: fuerst@uidaho.edu
Funding Source: Arnold and Mabel Beckman Foundation
External Link to Project Information:
Project Location: Moscow, ID

